Drug Response Prediction Based on Graph Topology Attention Network
-
摘要: 药物响应预测是生物医学研究中的重要课题,对推动癌症个性化治疗具有重要意义。尽管目前已有方法在药物响应预测方面取得了一定进展,然而,细胞系多组学数据的有效整合与药物特征的高效提取仍是当前研究面临的关键挑战。针对这一问题,本文提出一种基于图拓扑注意力网络的药物分子特征提取方法,并使用注意力机制融合多组学特征,进而实现药物响应预测。实验结果表明,本文所提出的模型在CCLE和GDSC两个数据集上均优于现有主流方法,消融实验进一步验证了模型结构与特征提取策略在本任务中的有效性。Abstract:
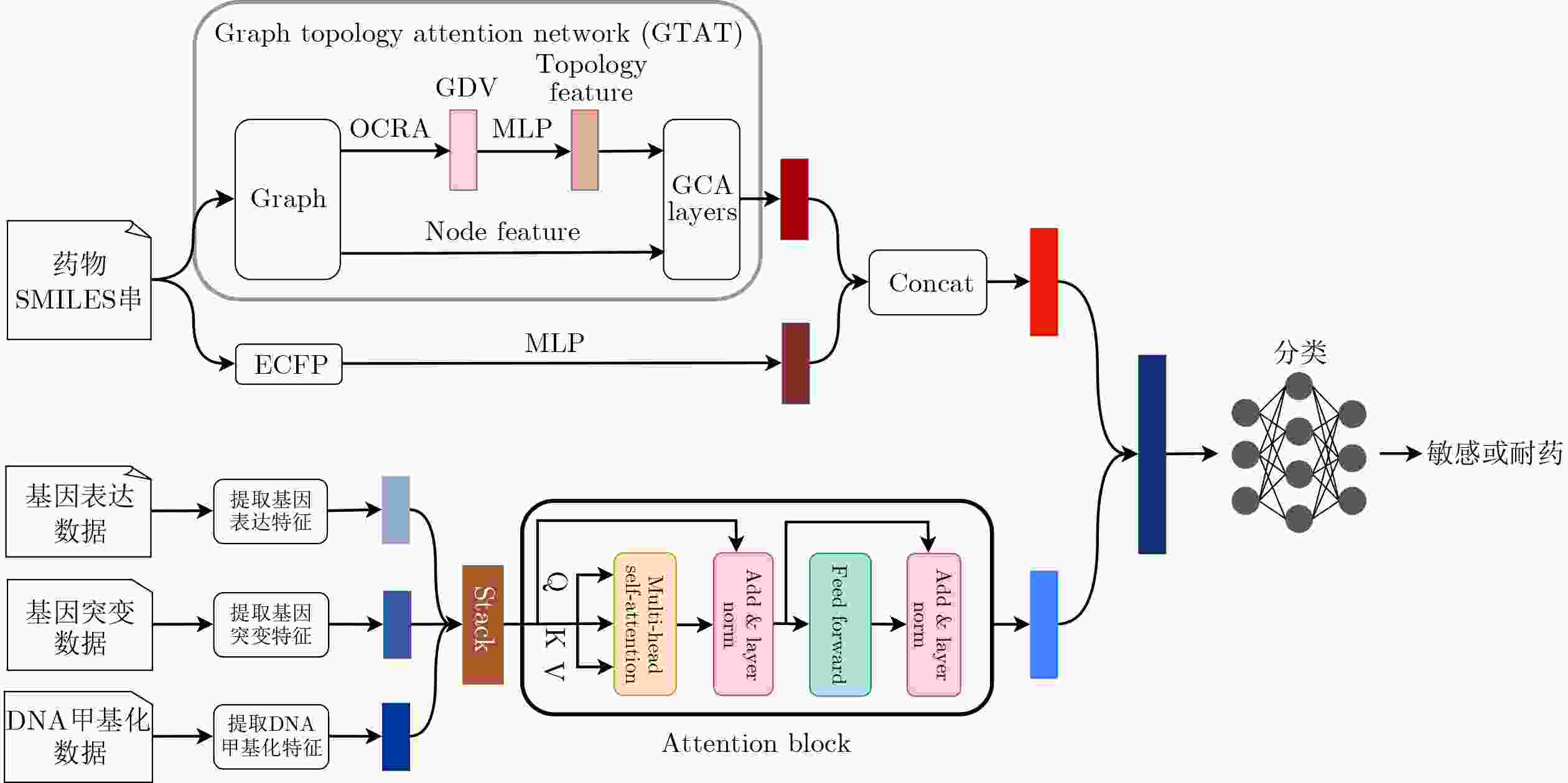
Objective A core goal in modern cancer research is to figure out why patients respond differently to the same therapy. Achieving this requires developing computational tools that combine genetic information and drug properties to forecast treatment outcomes, which is essential for advancing personalized oncology. Although some existing methods have made progress in predicting cancer drug responses, effectively extracting features of drugs and integrating multi-omics data from cell lines have become challenges. To address these challenges, employing Graph Neural Networks (GNNs) to process drug molecular graphs has become a promising strategy. This research proposes a model that utilizes a graph topology attention network to capture features from drug molecular graphs, while an attention mechanism is applied to integrate multi-omics data. Methods In this study, a drug response prediction method based on Graph Topology Attention Network(GTAT) is proposed. The model integrates topological graph information to predict drug responses in cell lines. The model utilizes drug SMILES strings to generate two distinct drug representations and incorporates multi-omics data for cell line characterization ( Fig. 1 ). For drug feature extraction, SMILES strings are first parsed to construct molecular graphs, which are then processed by the GTAT. This network captures both the topological information of the molecular graph-level and atom-level features, thereby producing structured molecular representations. Simultaneously, Extended Connectivity Fingerprints are computed from the same SMILES strings and transformed into continuous feature vectors via a Multi-Layer Perceptron (MLP). The graph-based drug representation and the fingerprint-based representation are subsequently concatenated to form a comprehensive drug feature vector. For cell line representation, multi-omics data are processed through omics-specific neural networks. The resulting features are fused using multi-head self-attention mechanisms, enabling the model to capture contextual interactions across omics modalities and generate an integrated cell line representation. Finally, the drug and cell line features are combined and fed into an MLP classifier to predict drug response outcomes. The proposed model effectively integrates heterogeneous biological data sources and significantly enhances prediction accuracy through multi-modal learning and attention-based feature fusion.Results and Discussions The proposed method achieves competitive performance on both GDSC and CCLE benchmark datasets ( Table 2 ). Specifically, on the GDSC dataset, our approach outperforms all competing methods across all four metrics—AUC, AUPR, F1-score, and Accuracy. Notably, it improves the AUPR by approximately 1.92% over the second-best method, MOFGCN, demonstrating its advantage in handling class imbalance. On the CCLE dataset, our method still achieves the best performance in terms of AUC and Accuracy. Although it is marginally lower than GADRP in AUPR and F1-score, the gap is minimal, and our approach exhibits more robust overall discriminative ability (as reflected by AUC). These results collectively validate the effectiveness and strong generalizability of our method in drug sensitivity prediction tasks. The observed variation in AUPR and F1-score performance between datasets can be attributed to inherent differences in sample size and class distribution characteristics. The limited scale of the CCLE dataset, combined with its specific class imbalance (approximately 4:1 ratio of resistant to sensitive samples), may constrain the model's capacity to fully learn the underlying data distribution, particularly for minority classes. In contrast, the GDSC dataset exhibits greater heterogeneity and a more pronounced class imbalance (approximately 8:1), which collectively contribute to increased prediction difficulty and consequently lower performance on certain metrics.Conclusions Accurately predicting drug response in cell lines remains a central challenge in precision medicine, with significant implications for accelerating drug development and advancing personalized treatment. However, constructing a high-accuracy predictive model capable of effectively integrating multi-source biological information is difficult due to the complexity of drug molecular structures and inherent heterogeneity of cell lines. To address this, a cell line drug response prediction model based on Graph Topology Attention Network is proposed. This model employs the graph topology attention network to extract molecular graph features of drugs, which are then fused with molecular fingerprint features. Meanwhile, multi-omics features of cell lines are integrated using an attention mechanism. Experimental results demonstrate that the proposed model achieves superior performance over existing state-of-the-art benchmarks on the employed dataset. This study provides a new perspective for predicting cell line drug response. Certain limitations are acknowledged, such as the use of only three types of omics features for cell line representation and the influence of sample size on predictive outcomes. The integration of more diverse omics features, the application of pre-trained large-scale models, and the clinical translation for personalized medicine will be the primary focus of future work. -
表 1 实验数据集统计信息
数据集 细胞系数量 药物数量 细胞系药物响应数量 GDSC 561 222 100572 CCLE 317 24 7307 表 2 在GDSC数据集和CCLE数据集上的性能比较
方法 GDSC CCLE AUC AUPR F1-score Accuracy AUC AUPR F1-score Accuracy DeepCDR 0.8240 0.4853 0.4746 0.8754 0.9546 0.8639 0.8224 0.9324 MOFGCN 0.8327 0.4945 0.4861 0.8764 0.9613 0.8893 0.8204 0.9291 GraphCDR 0.8296 0.4869 0.4807 0.8741 0.9483 0.8746 0.8253 0.9322 GADRP 0.8220 0.4792 0.4709 0.8743 0.9628 0.8994 0.8347 0.9365 Ours 0.8383 0.5040 0.4864 0.8783 0.9635 0.8916 0.8304 0.9405 注:最好结果和次好结果分别用粗体和下划线突出显示 表 3 模型消融实验
方法 AUC AUPR F1-score Accuracy w/o GE 0.8340 0.4976 0.4792 0.8708 w/o GM 0.8372 0.5010 0.4833 0.8748 w/o DM 0.8324 0.4916 0.4745 0.8743 w/o Graph 0.8265 0.4847 0.4713 0.8752 w/o ECFP 0.7737 0.3458 0.3978 0.8263 w linear 0.8250 0.4844 0.4683 0.8734 w mean 0.8344 0.5033 0.4804 0.8763 w max 0.8302 0.4950 0.4773 0.8727 Ours 0.8383 0.5040 0.4864 0.8783 注:最好结果用粗体突出显示 表 4 预测对两种药物敏感的前八个癌细胞系
Drug Rank Cancer cell line PMID or DOI Dasatinib 1 A-375 10.1016 /j.tetlet.2024.1553652 HT-144 18823558 3 MEL-JUSO N/A 4 BFTC-905 32370101 5 Hep 3B2.1-7 N/A 6 HSC-3 N/A 7 RKO 40481178 8 SK-MEL-30 N/A GSK690693 1 RPMI- 8402 N/A 2 NALM-6 19064730 3 SU-DHL-6 39632683 4 ALL-SIL N/A 5 SIG-M5 N/A 6 697 19064730 7 RCH-ACV 19064730 8 SU-DHL-5 39632683 -
[1] BRAY F, LAVERSANNE M, SUNG H, et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA: A Cancer Journal for Clinicians, 2024, 74(3): 229–263. doi: 10.3322/caac.21834. [2] 周涛, 叶鑫宇, 刘凤珍, 等. 利用跨模态轻量级YOLOv5模型的PET/CT肺部肿瘤检测[J]. 电子与信息学报, 2024, 46(2): 624–632. doi: 10.11999/JEIT230052.ZHOU Tao, YE Xinyu, LIU Fengzhen, et al. CL-YOLOv5: PET/CT lung cancer detection with cross-modal lightweight YOLOv5 model[J]. Journal of Electronics & Information Technology, 2024, 46(2): 624–632. doi: 10.11999/JEIT230052. [3] 鲍振申, 李先彬, 刘晓晖, 等. 基于网络最短路径的miRNA对疾病异常细胞功能影响的量化方法[J]. 广州大学学报: 自然科学版, 2025, 24(6): 69–77.BAO Zhenshen, LI Xianbin, LIU Xiaohui, et al. A computational method for quantifying the impacts on abnormal disease cellular functions of miRNAs based on the shortest path of the network[J]. Journal of Guangzhou University: Natural Science Edition, 2025, 24(6): 69–77. [4] KURU H I, CICEK A E, and TASTAN O. From cell lines to cancer patients: Personalized drug synergy prediction[J]. Bioinformatics, 2024, 40(5): btae134. doi: 10.1093/bioinformatics/btae134. [5] 苏树智, 张开宇, 王子莹, 等. 基于跨视角相似度顺序保持的基因特征提取方法[J]. 电子与信息学报, 2023, 45(1): 317–324. doi: 10.11999/JEIT211126.SU Shuzhi, ZHANG Kaiyu, WANG Ziying, et al. A gene feature extraction method based on across-view similarity order preserving[J]. Journal of Electronics & Information Technology, 2023, 45(1): 317–324. doi: 10.11999/JEIT211126. [6] YANG Wanjuan, SOARES J, GRENINGER P, et al. Genomics of Drug Sensitivity in Cancer (GDSC): A resource for therapeutic biomarker discovery in cancer cells[J]. Nucleic Acids Research, 2013, 41(D1): D955–D961. doi: 10.1093/nar/gks1111. [7] 许鹏, 谢斌, 鲍振申, 等. 基于自编码器的疾病相关miRNAs的预测方法[J]. 广州大学学报: 自然科学版, 2024, 23(1): 12–19. doi: 10.3969/j.issn.1671-4229.2024.01.002.XU Peng, XIE Bin, BAO Zhenshen, et al. Predictive method for disease-associated miRNAs based on autoencoders[J]. Journal of Guangzhou University: Natural Science Edition, 2024, 23(1): 12–19. doi: 10.3969/j.issn.1671-4229.2024.01.002. [8] 顾丽丽, 郑泽昆, 方紫婵, 等. 基于miRNA-gene调控网络的miRNA重要性评估[J]. 广州大学学报: 自然科学版, 2024, 23(3): 8–14. doi: 10.3969/j.issn.1671-4229.2024.03.002.GU Lili, ZHENG Zekun, FANG Zichan, et al. Evaluation of the importance of miRNAs based on miRNA-gene regulatory networks[J]. Journal of Guangzhou University: Natural Science Edition, 2024, 23(3): 8–14. doi: 10.3969/j.issn.1671-4229.2024.03.002. [9] 阮东升, 施哲彬, 王嘉辉, 等. 结合跨模态特征激励与双分支交叉注意力融合的左心房疤痕分割方法[J]. 电子与信息学报, 2025, 47(5): 1596–1608. doi: 10.11999/JEIT240775.RUAN Dongsheng, SHI Zhebin, WANG Jiahui, et al. Left atrial scar segmentation method combining cross-modal feature excitation and dual branch cross attention fusion[J]. Journal of Electronics & Information Technology, 2025, 47(5): 1596–1608. doi: 10.11999/JEIT240775. [10] 张淑芳, 李予辉, 李炳志. DNA存储场景下基于引物索引矩阵的文件高效随机检索方法[J]. 电子与信息学报, 2024, 46(6): 2568–2577. doi: 10.11999/JEIT230717.ZHANG Shufang, LI Yuhui, and LI Bingzhi. Efficient file random access method based on primer index matrix in DNA storage scenarios[J]. Journal of Electronics & Information Technology, 2024, 46(6): 2568–2577. doi: 10.11999/JEIT230717. [11] GUAN Nana, ZHAO Yan, WANG Chunchun, et al. Anticancer drug response prediction in cell lines using weighted graph regularized matrix factorization[J]. Molecular Therapy - Nucleic Acids, 2019, 17: 164–174. doi: 10.1016/j.omtn.2019.05.017. [12] WANG Yongcui, FANG Jianwen, and CHEN Shilong. Inferences of drug responses in cancer cells from cancer genomic features and compound chemical and therapeutic properties[J]. Scientific Reports, 2016, 6: 32679. doi: 10.1038/srep32679. [13] WAN Qian and PAL R. An ensemble based top performing approach for NCI-DREAM drug sensitivity prediction challenge[J]. PLoS One, 2014, 9(6): e101183. doi: 10.1371/journal.pone.0101183. [14] ZHONG Jiancheng, ZOU Zhiwei, QIU Jie, et al. ScFold: A GNN-based model for efficient inverse folding of short-chain proteins via spatial reduction[J]. Briefings in Bioinformatics, 2025, 26(2): bbaf156. doi: 10.1093/bib/bbaf156. [15] 顾丽丽, 蒋静梅, 王慧静, 等. 基于多模态图神经网络的miRNA-疾病关联预测模型[J]. 广州大学学报: 自然科学版, 2025, 24(6): 78–88.GU Lili, JIANG Jingmei, WANG Huijing, et al. A multimodal graph neural network model for miRNA-disease association prediction[J]. Journal of Guangzhou University: Natural Science Edition, 2025, 24(6): 78–88. [16] LIU Xuan, SONG Congzhi, HUANG Feng, et al. GraphCDR: A graph neural network method with contrastive learning for cancer drug response prediction[J]. Briefings in Bioinformatics, 2022, 23(1): bbab457. doi: 10.1093/bib/bbab457. [17] PENG Wei, LIU Hancheng, DAI Wei, et al. Predicting cancer drug response using parallel heterogeneous graph convolutional networks with neighborhood interactions[J]. Bioinformatics, 2022, 38(19): 4546–4553. doi: 10.1093/bioinformatics/btac574. [18] YANG Peisheng, HE Changxiang, ZHANG Ping, et al. MHGCN: A multi-channel hybrid graph convolutional neural network for cancer drug response prediction[J]. IEEE Transactions on Computational Biology and Bioinformatics, 2025, 22(5): 2241–2251. doi: 10.1109/TCBBIO.2025.3594056. [19] LIU Qiao, HU Zhiqiang, JIANG Rui, et al. DeepCDR: A hybrid graph convolutional network for predicting cancer drug response[J]. Bioinformatics, 2020, 36(S2): i911–i918. doi: 10.1093/bioinformatics/btaa822. [20] KIM S, CHEN Jie, CHENG Tiejun, et al. PubChem 2019 update: Improved access to chemical data[J]. Nucleic Acids Research, 2019, 47(D1): D1102–D1109. doi: 10.1093/nar/gky1033. [21] RAMSUNDAR B, EASTMAN P, WALTERS P, et al. Deep Learning for the Life Sciences: Applying Deep Learning to Genomics, Microscopy, Drug Discovery, and More[M]. Sebastopol: O'Reilly Media, 2019. (查阅网上资料, 请补充引用页码). [22] IORIO F, KNIJNENBURG T A, VIS D J, et al. A landscape of pharmacogenomic interactions in cancer[J]. Cell, 2016, 166(3): 740–754. doi: 10.1016/j.cell.2016.06.017. [23] DONG Zuoli, ZHANG Naiqian, LI Chun, et al. Anticancer drug sensitivity prediction in cell lines from baseline gene expression through recursive feature selection[J]. BMC Cancer, 2015, 15(1): 489. doi: 10.1186/s12885-015-1492-6. [24] SHEN Jiahao, AIN Q T, LIU Yaohua, et al. GTAT: Empowering graph neural networks with cross attention[J]. Scientific Reports, 2025, 15(1): 4760. doi: 10.1038/s41598-025-88993-3. [25] VASWANI A, SHAZEER N, PARMAR N, et al. Attention is all you need[C]. The 31st Conference on Neural Information Processing Systems, Long Beach, USA, 2017: 6000–6010. [26] MILENKOVIĆ T, NG W L, HAYES W, et al. Optimal network alignment with graphlet degree vectors[J]. Cancer Informatics, 2010, 9: 121–137. doi: 10.4137/cin.s4744. [27] HOČEVAR T and DEMŠAR J. A combinatorial approach to graphlet counting[J]. Bioinformatics, 2014, 30(4): 559–565. doi: 10.1093/bioinformatics/btt717. [28] VELIČKOVIĆ P, CUCURULL G, CASANOVA A, et al. Graph attention networks[C]. The 6th International Conference on Learning Representations, Vancouver, Canada, 2018. [29] WILLETT P. The calculation of molecular structural similarity: Principles and practice[J]. Molecular Informatics, 2014, 33(6/7): 403–413. doi: 10.1002/minf.201400024. [30] REYAD M, SARHAN A M, and ARAFA M. A modified Adam algorithm for deep neural network optimization[J]. Neural Computing and Applications, 2023, 35(23): 17095–17112. doi: 10.1007/s00521-023-08568-z. [31] PENG Wei, CHEN Tielin, and DAI Wei. Predicting drug response based on multi-omics fusion and graph convolution[J]. IEEE Journal of Biomedical and Health Informatics, 2022, 26(3): 1384–1393. doi: 10.1109/JBHI.2021.3102186. [32] WANG Hong, DAI Chong, WEN Yuqi, et al. GADRP: Graph convolutional networks and autoencoders for cancer drug response prediction[J]. Briefings in Bioinformatics, 2023, 24(1): bbac501. doi: 10.1093/bib/bbac501. [33] DARWISH S, DAVANI-DAVARI D, TONG S, et al. Synthesis and evaluation of cyclic peptide-dasatinib conjugates as anti-melanoma agents[J]. Tetrahedron Letters, 2024, 152: 155365. doi: 10.1016/j.tetlet.2024.155365. [34] EUSTACE A J, CROWN J, CLYNES M, et al. Preclinical evaluation of dasatinib, a potent Src kinase inhibitor, in melanoma cell lines[J]. Journal of Translational Medicine, 2008, 6(1): 53. doi: 10.1186/1479-5876-6-53. [35] LEVY D S, KAHANA J A, and KUMAR R. AKT inhibitor, GSK690693, induces growth inhibition and apoptosis in acute lymphoblastic leukemia cell lines[J]. Blood, 2009, 113(8): 1723–1729. doi: 10.1182/blood-2008-02-137737. -




 下载:
下载:

 下载:
下载:
